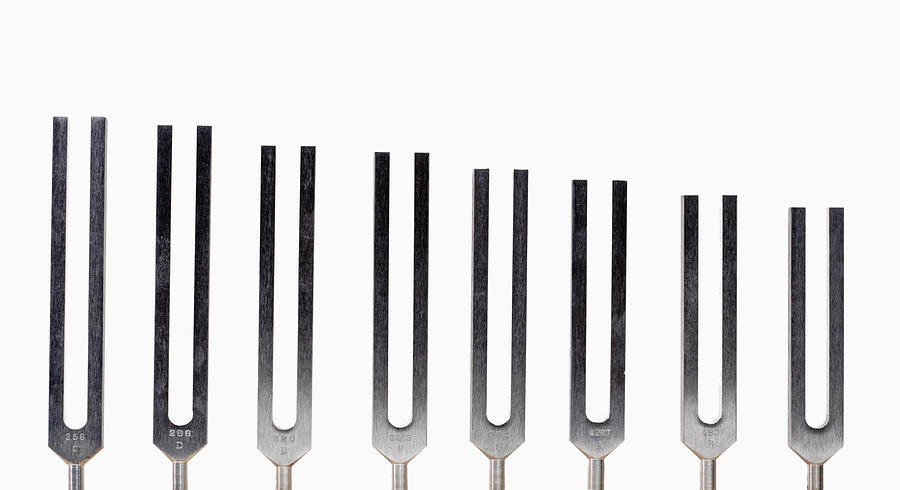

This bacterium harbors a main chromosome (Chr1) of 2.96 Mbp and a 1.07-Mbp secondary replicon (Chr2). In previous works, we tackled this issue in Vibrio cholerae, the causative agent of cholera disease. 1a, right) when the transcriptional activity and ribosome numbers increase by 10- and 15-fold, respectively. Thus, the dosage and expression of the aforementioned genes peak during exponential growth phase (Fig. Therefore, it was proposed that the oriC-proximal location of ribosomal and transcription genes allows the recruitment of multifork replication for growth optimization purposes. This leads to replication-associated gene dosage gradients along the ori-ter axis during exponential growth (Fig. Consequently, chromosomes trigger replication more than once before cytokinesis, overlapping successive DNA duplication rounds, a phenomenon called multifork replication (Fig. These microorganisms divide faster than the time required for genome duplication. In fast-growing bacteria, the genes coding for transcription and translation machineries locate near the oriC. Notable examples are genes encoding the flux of the genetic information. In addition, recent studies have showcased an increasing number of traits whose expression is influenced by the genomic position of its encoding genes. Remarkably, key genes coding for nucleoid-associated proteins, RNA polymerase modulators, topoisomerases, and energy production are arranged along the ori-ter axis following the temporal order of their expression during growth phases. Recent studies indicate that gene order within the chromosome may play a relevant role in harmonizing the genome structure with cell physiology. Large inversions occur preferentially symmetrically with respect to the ori-ter axis to avoid the emergence of replichore size imbalance. For instance, essential genes are overrepresented in the replicative leading strand to avoid head-on collisions between the replication and transcription machineries. 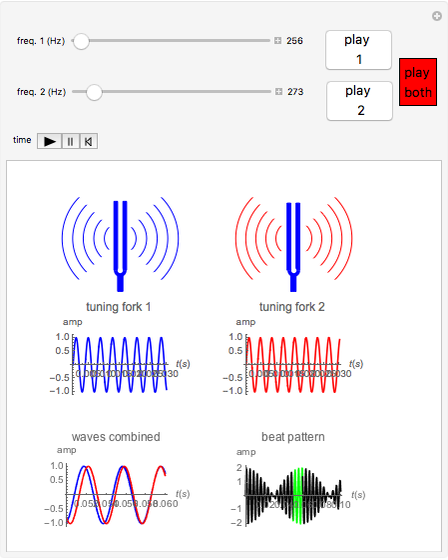
This organizes the genome along an ori-ter axis that interplays with cell physiology (Fig. Replication begins at a sole replication origin ( oriC), proceeding bidirectionally along two equally sized replichores until the terminal region ( ter). Bacterial chromosomes are highly variable in their gene content, but highly conserved in terms of the order of core genes in the chromosomes. 
Their relative simplicity and the increasing amount of available data render bacterial genomes ideal models to study this subject. Genome structure may contribute to integrate these many simultaneous processes occurring on the same template. Replication, gene expression, and segregation are tightly coordinated with the cell cycle to preserve cellular homeostasis. Hence, this could be another mechanism coordinating DNA replication to bacterial growth. However, besides of its essential function in translation, their genomic position sustains an optimal macromolecular crowding essential for maximizing growth. The genomic location of RP genes ensures its optimal dosage. In hyperosmotic conditions, when crowding differences are minimized, the growth rate and replication dynamics were highly alleviated in these strains. Accordingly, cytoplasm fluidity was higher in mutants where S10 is most distant from oriC. Since RP constitutes a large proportion of cell mass, lower S10 dosage could lead to changes in macromolecular crowding, impacting cell physiology. Such changes increased as a function of oriC-S10 distance. Deep sequencing revealed that S10 relocation altered chromosomal replication dynamics and genome-wide transcription. Strikingly, we observed that in Vibrio cholerae, protein production capacity was independent of S10 position. We hypothesized that S10 dosage perturbations impact protein synthesis capacity. However, a mechanism linking S10 dosage to cell physiology has still not been determined. Relocation of s10-spc-α locus (S10), which codes for most of the RP, to ectopic genomic positions shows that its relative distance to the oriC correlates to a reduction on its dosage, its expression, and bacterial growth rate. This trait allows optimizing their expression during exponential phase since oriC neighboring regions are in higher dose due to multifork replication. In fast-growing bacteria, the genomic location of ribosomal protein (RP) genes is biased towards the replication origin ( oriC).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
